


Next: 3 Downloading and loading
Up: Inferring patterns of migration
Previous: 1 Introduction
Contents
We consider
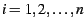 subpopulation. The available
data from which the inferences are to be drawn are vectors
of gene (allele) frequencies in these subpopulations, at a
number of different loci. Differences between local gene
frequencies is assumed to be a result of local genetic drift
balanced by migration (or mutation). The pattern of
migration between these subpopulation is described by a
migration matrix
subpopulation. The available
data from which the inferences are to be drawn are vectors
of gene (allele) frequencies in these subpopulations, at a
number of different loci. Differences between local gene
frequencies is assumed to be a result of local genetic drift
balanced by migration (or mutation). The pattern of
migration between these subpopulation is described by a
migration matrix
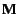 with elements
with elements 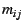 denoting the
probability, or the proportion, of genes which originates
from subpopulation
denoting the
probability, or the proportion, of genes which originates
from subpopulation 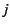 , given migration to subpopulation
, given migration to subpopulation
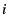 . The objective of analysis is to obtain an estimate of
the migration matrix
. The objective of analysis is to obtain an estimate of
the migration matrix
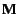 . In general, we will be
interested in migration matrices of certain biologically
interesting forms only, involving a small number of
parameters, which we hope to be able to estimate. The
effective size, or at least, the relative size, of each
subpopulation,
. In general, we will be
interested in migration matrices of certain biologically
interesting forms only, involving a small number of
parameters, which we hope to be able to estimate. The
effective size, or at least, the relative size, of each
subpopulation, 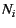 , is treated as known parameters. All
subpopulations are also assumed to receive a small amount of
immigrant genes from a large `outside world' population with
gene frequency
, is treated as known parameters. All
subpopulations are also assumed to receive a small amount of
immigrant genes from a large `outside world' population with
gene frequency 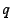 .
.
With these assumptions, one can show that the gene
frequencies, at equilibrium, provided that the fluctuations
around 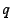 are small, have covariance matrix
are small, have covariance matrix
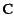 ,
satisfying the matrix equation
,
satisfying the matrix equation
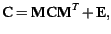 |
(1) |
where
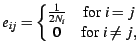 |
(2) |
Using the solution of (1), the idea is to compute
the likelihood of the data approximately by assuming
multivariate normality. It is then possible to obtain
maximum likelihood estimates of the parameters of any model
for the migration matrix using numerical methods.



Next: 3 Downloading and loading
Up: Inferring patterns of migration
Previous: 1 Introduction
Contents
Jarle Tufto
2001-08-28
![]() are small, have covariance matrix
are small, have covariance matrix
![]() ,
satisfying the matrix equation
,
satisfying the matrix equation